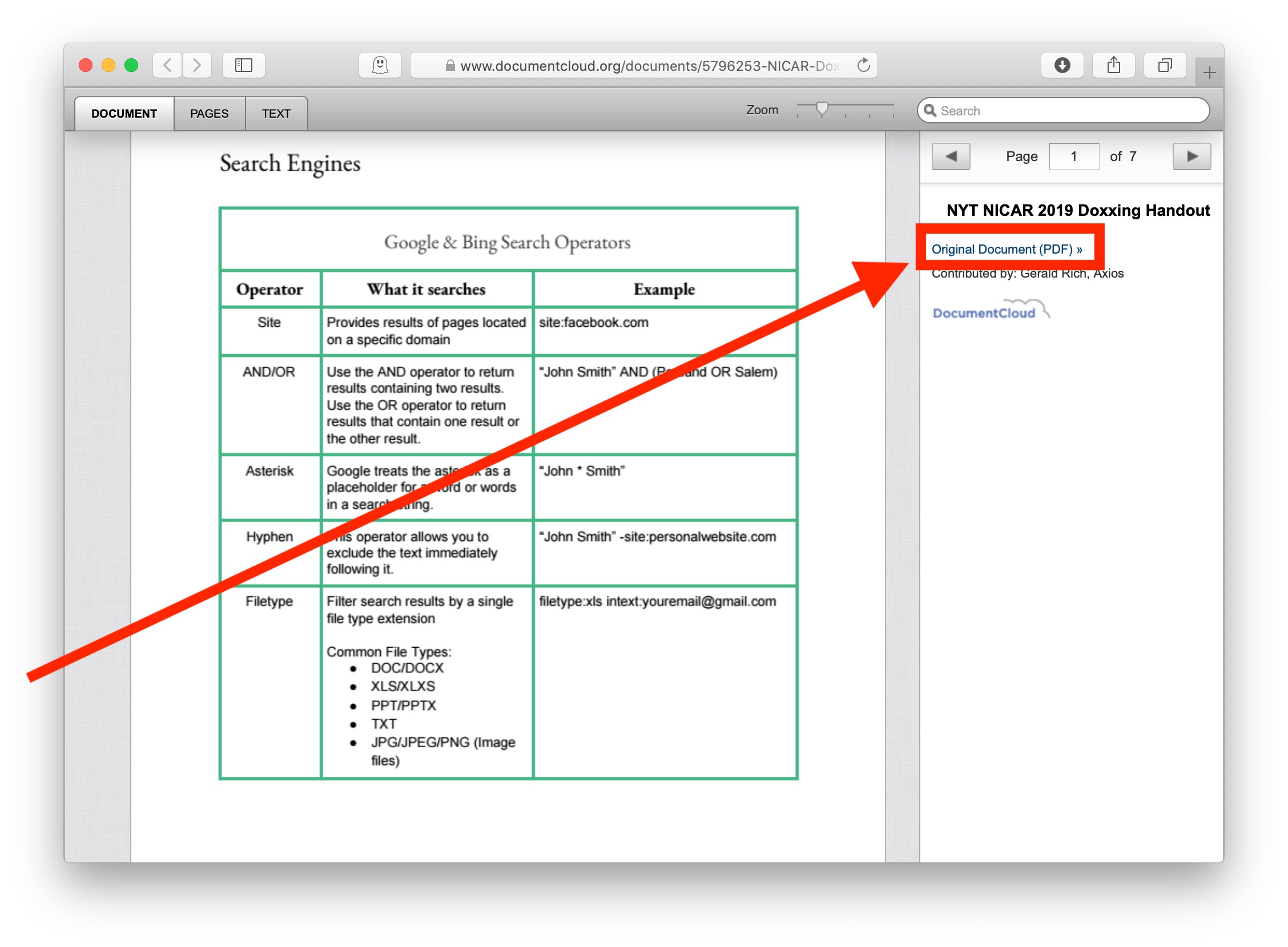
Does anyone have an explanation for this output? In the example pictures i have zoomed in on an example of this with coeluting peaks where one peak is larger than the other in AMDIS and chemstation while its the opposite in MzMine. 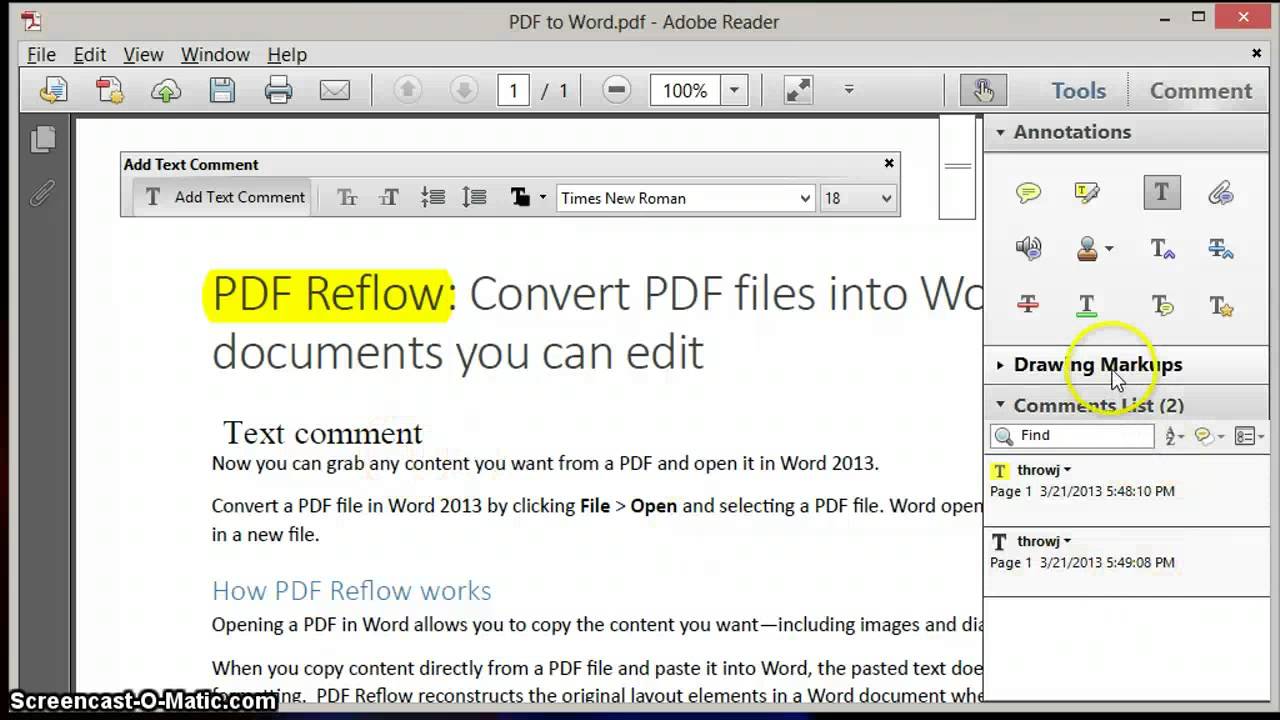
The differences in the chromatogram are minor, if you take an overall look the chromatogram looks similar but when you look close there's differences. But this is not the case for me as you can see in the attached pictures. According to my knowledge, the CDF file should create a similar chromatogram out of the CDF file in both programs. However, when I open the CDF file with AMDIS the chromatogram looks similar to that of chemstation. In order to open them with MzMine i need to convert them to CDF files. D files to CDF.D is the custom format I get when the samples have been run on our instrument and these files can be opened with chemstation. First I thought that the issue might be the conversion from. When I open the CDF files from one of my GC-MS runs in MzMine the TIC looks different in MzMine compared to AMDIS and chemstation. I've had some issues with the total ion chromatogram (TIC) when importing CDF files into MzMine.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
